Antibody characterization process
Introduction
Many of the assays used in the ENCODE project involve the use of antibodies. Choosing an appropriate antibody involves a regimented and extensive antibody characterization process to establish a line of evidence that these antibodies specifically and accurately bind the target in question. It has long been a goal of the project to make this process and its results transparent.
Basic organization | Data from previous and external projects
Antibody Lot Eligibility Statuses | Antibody Characterization Statuses | Antibody Standards
Search | How To
Basic organization
The basic unit of organization is the antibody lot, which is defined as a unique lot-productID-source combination. Each antibody lot has a unique ENCODE accession. Unless an antibody is against a histone modification, it is required by the current ENCODE project standards to be characterized in each cell type and species (see section on Standards documents). Each antibody lot will have multiple supporting characterizations, which can include dot blot assays, immunoblots, and/or mass spectrometry results. Each characterization submitted in support of an antibody lot’s specificity to the intended target is reviewed against the current standards document and the result is reflected in its status. For each antibody lot, its eligibility for use in the current project is determined by having characterizations that comply to the current standards for the cell/tissue type and species being investigated.
Data from previous and external projects
The ENCODE portal not only has data from the current ENCODE project, but also data from the previous ENCODE projects and some data from modENCODE and Epigenomics ROADMAP. Standards are always evolving and improving. Therefore, as these projects did not have the same structure or current standards as our current project, any antibody information and characterizations that we did import are marked as not reviewed. This is not meant to imply anything about the quality of the antibody or characterization other than it is not in the scope of the current ENCODE project to review these imported characterizations and that we will not use any of these lots on current incoming data. However, if a current ENCODE submitting lab would like to use a previously imported lot, they will ensure the supporting characterizations conform to the current standards and request a review.
Antibody characterization status
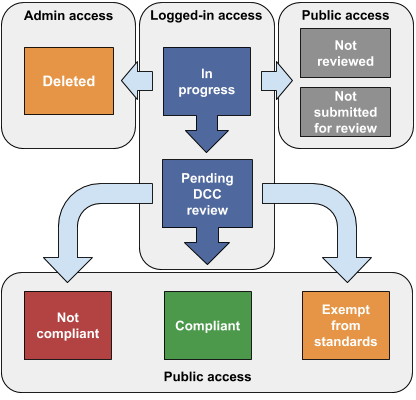
- compliant : The characterization has been found to be compliant with the attached standards document.
- exempt from standards: The characterization has been exempt from standards by the ENCODE Antibody Review Panel.
- not compliant : The characterization has been found to be not compliant with the attached standards document.
- in progress : This is an internal state and should not be seen on the public site. It indicates that this antibody is in consideration for use but its evidence is not ready for review at this time.
- pending dcc review : This characterization is completed by the submitting lab and waiting for the review process. Like "in progress," internal state and should not be seen on the public site.
- not reviewed : This characterization is from a previous or external project. It will likely have data associated with it, but it will not be reviewed by the current ENCODE project.
- not submitted for review : This characterization was either attempted or proposed but will not be submitted by the lab for the review process.
Antibody Lot eligibility status
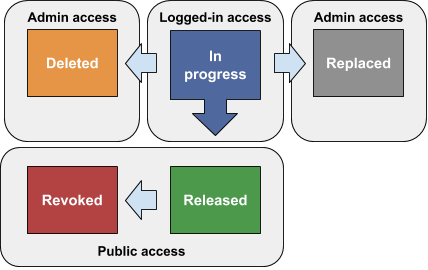
Standards documents
Characterization of antibodies is one process that the consortium places much rigor in standardizing, in order to determine whether an antibody lot reproducibly binds its intended target. Antibody lot characterization methods and standards will vary depending on the target of the antibody (e.g. histones, transcription factors). Similarly, as new methods are developed, the standards for antibody lot characterization will evolve. For these reasons, there will be multiple versions of the ENCODE antibody characterization standard documents produced over time. Explore the antibody characterization guidelines page for previous and current versions. An example of how standards documents are associated with compliant characterizations can be found in lower right of Figure 4 on this page.
Search page and facets
The antibody search page uses color-coding to indicate the eligibility status of the antibody lots. A lot may have differing eligibility based on tissue/cell type or species. There are facets to search the eligibility status, source, clonality, and host organisms of the antibody-lot and facets to select the lab and target organism of the related characterizations.
Antibody lot pages
Each antibody lot has a page with five sections: accession/title, eligibility, lot information, characterizations, related experiments. The header of the page includes the accession/title in grey, followed by the eligibility organized by cell type and species, followed by the lot information.
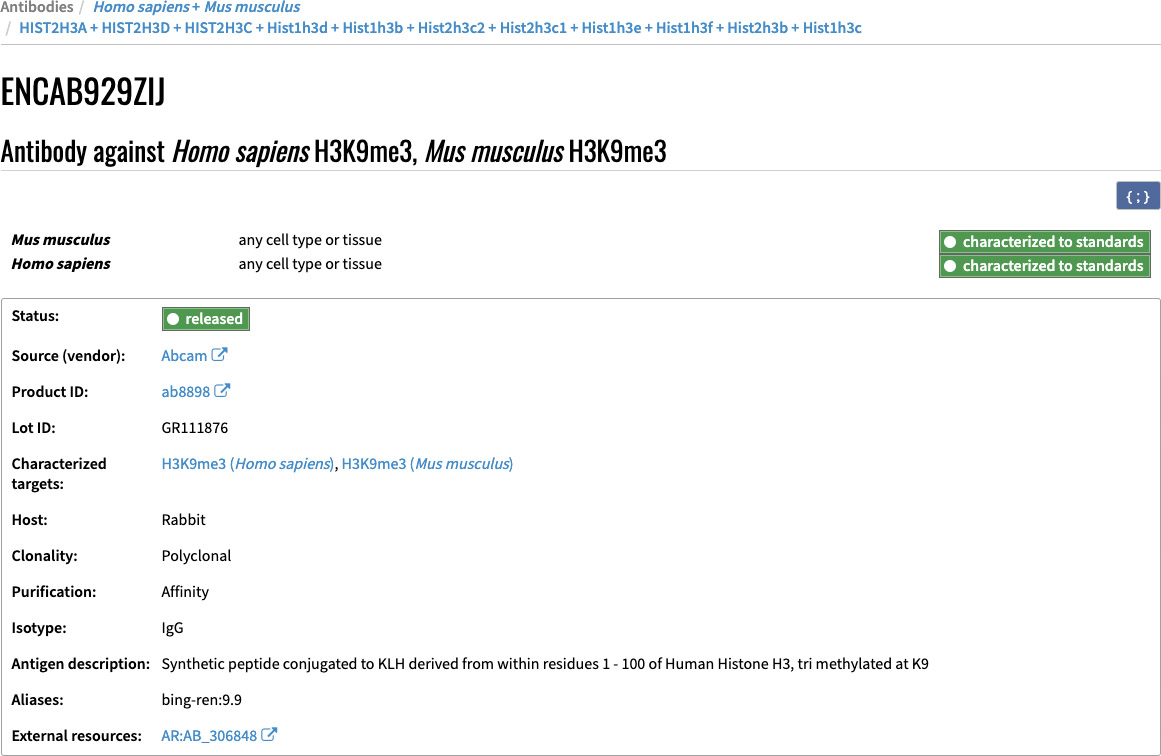
Figure 3. Header section of the antibody lot page including accession, eligibility information and lot details.
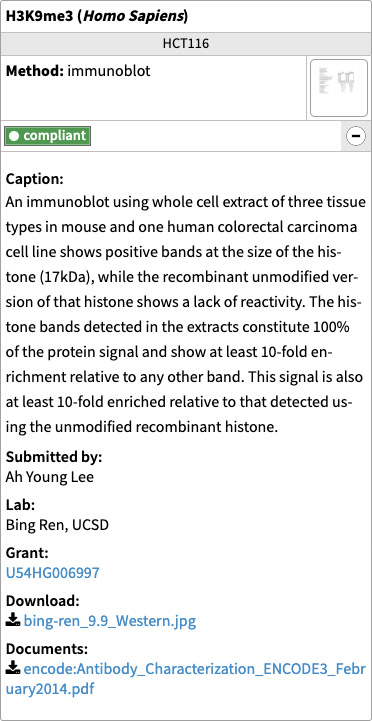
The characterization section has the related characterizations organized by species. Each characterization includes the documents (images, tables, pdfs) being used as evidence along with a caption. It also indicates its status, target species and submitter information. If the characterization is considered compliant, it will also have a link to the standards document it was reviewed against.
Figure 4. Example characterization block found on the antibody lot page.
All of the experiments using a particular antibody lot can be accessed at the bottom of the antibody lot page. The first five are listed directly on the page as in the figure. The View All link will bring up a search page of all related experiments.
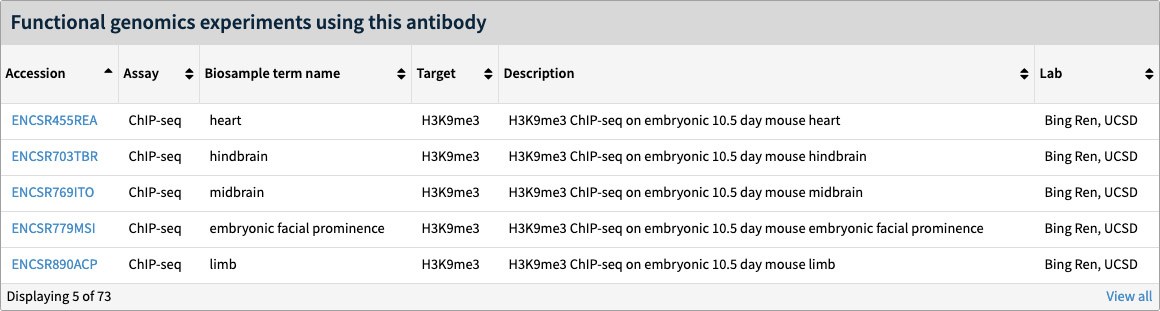
Figure 5. Related experiments section of the antibody lot page.
Updated April 2018
