ENCAB426WVA
Antibody against Homo sapiens ZMYM3
Homo sapiens
K562, GM12878, HepG2, MCF-7
characterized to standards with exemption
Homo sapiens
any cell type or tissue
partially characterized
- Status
- released
- Source (vendor)
- CDI
- Product ID
- JH39.2.2F10
- Lot ID
- HAIB-001
- Characterized targets
- ZMYM3 (Homo sapiens)
- Host
- mouse
- Clonality
- monoclonal
- Purification
- Protein G
- Isotype
- IgG2a
- Antigen description
- Monospecific for human ZMYM3
- External resources
Characterizations
ZMYM3 (Homo sapiens)
not compliant
- Caption
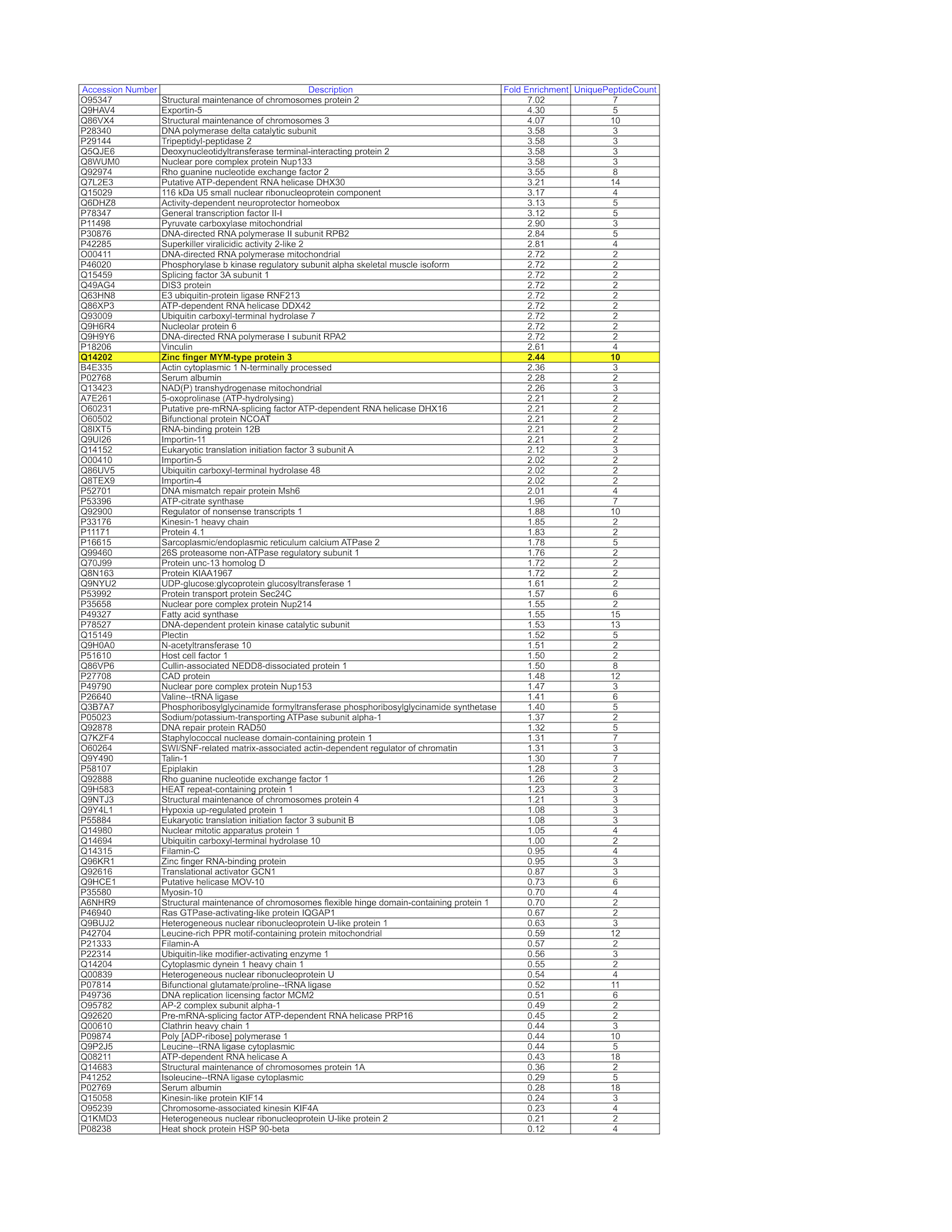
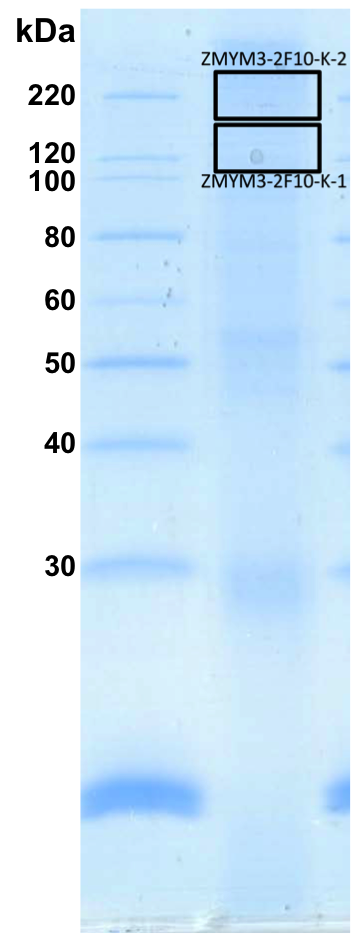
- K562 whole cell lysate was immunoprecipitated using the primary antibody (CDI; JH39.2.2F10). The IP fraction was loaded on a 12% Bio-Rad TGX gel and separated with the Bio-Rad Tetra Cell system. Gel fragments (rectangle outline) corresponding to the bands indicated on the Coomassie Blue stained gel image were excised and sent to the University of Alabama at Birmingham Cancer Center Mass Spectrometry/Proteomics Shared Facility. Analysis of gel fragment 1 from K562: The sample was analyzed on a LTQ XL Linear Ion Trap Mass Spectrometer by LC-ESI-MS/MS. Peptides were identified using SEQUEST tandem mass spectral analysis with probability based matching at p < 0.05. SEQUEST results were reported with ProteinProphet protXML Viewer (TPP v4.4 JETSTREAM) and filtered for a minimum probability of 0.9. All protein hits that met these criteria were reported, including common contaminants. Fold enrichment for each protein reported was determined using a custom script based on the FC-B score calculation from the reference Mellacheruvu et al., 2013. The CRAPome: a contaminant repository for affinity purification mass spectrometry data. Nat. Methods. 10(8):730-736. Doi:10.1038/nmeth.2557. The target protein, ZMYM3, was identified as the 26th enriched protein and the 2nd transcription factor based on IP-Mass Spectrometry.
- Reviewer comment
- Factor of interest is not within the top 20 proteins ranked by fold enrichment
- Submitted by
- Mark Mackiewicz
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998
- Download
- ZMYM3-1.png
ZMYM3 (Homo sapiens)
compliant
- Caption
- K562 whole cell lysate was immunoprecipitated using the primary antibody (CDI; JH39.2.2F10). The IP fraction was loaded on a 12% Bio-Rad TGX gel and separated with the Bio-Rad Tetra Cell system. Gel fragments (rectangle outline) corresponding to the bands indicated on the Coomassie Blue stained gel image were excised and sent to the University of Alabama at Birmingham Cancer Center Mass Spectrometry/Proteomics Shared Facility. Analysis of gel fragment 2 from K562: The sample was analyzed on a LTQ XL Linear Ion Trap Mass Spectrometer by LC-ESI-MS/MS. Peptides were identified using SEQUEST tandem mass spectral analysis with probability based matching at p < 0.05. SEQUEST results were reported with ProteinProphet protXML Viewer (TPP v4.4 JETSTREAM) and filtered for a minimum probability of 0.9. All protein hits that met these criteria were reported, including common contaminants. Fold enrichment for each protein reported was determined using a custom script based on the FC-B score calculation from the reference Mellacheruvu et al., 2013. The CRAPome: a contaminant repository for affinity purification mass spectrometry data. Nat. Methods. 10(8):730-736. Doi:10.1038/nmeth.2557. The target protein, ZMYM3, was identified as the 1st enriched protein and the 1st transcription factor based on IP-Mass Spectrometry.
- Submitted by
- Mark Mackiewicz
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998
- Download
- ZMYM3-2.png
ZMYM3 (Homo sapiens)
not submitted for review by lab
- Reviewer comment
- IP gel submitted for mass spectrometry analysis
- Submitted by
- Mark Mackiewicz
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998
- Download
- ZMYM3_IP-Commassie1.png
ZMYM3 (Homo sapiens)
K562GM12878HepG2MCF-7
exempt from standards
- Caption
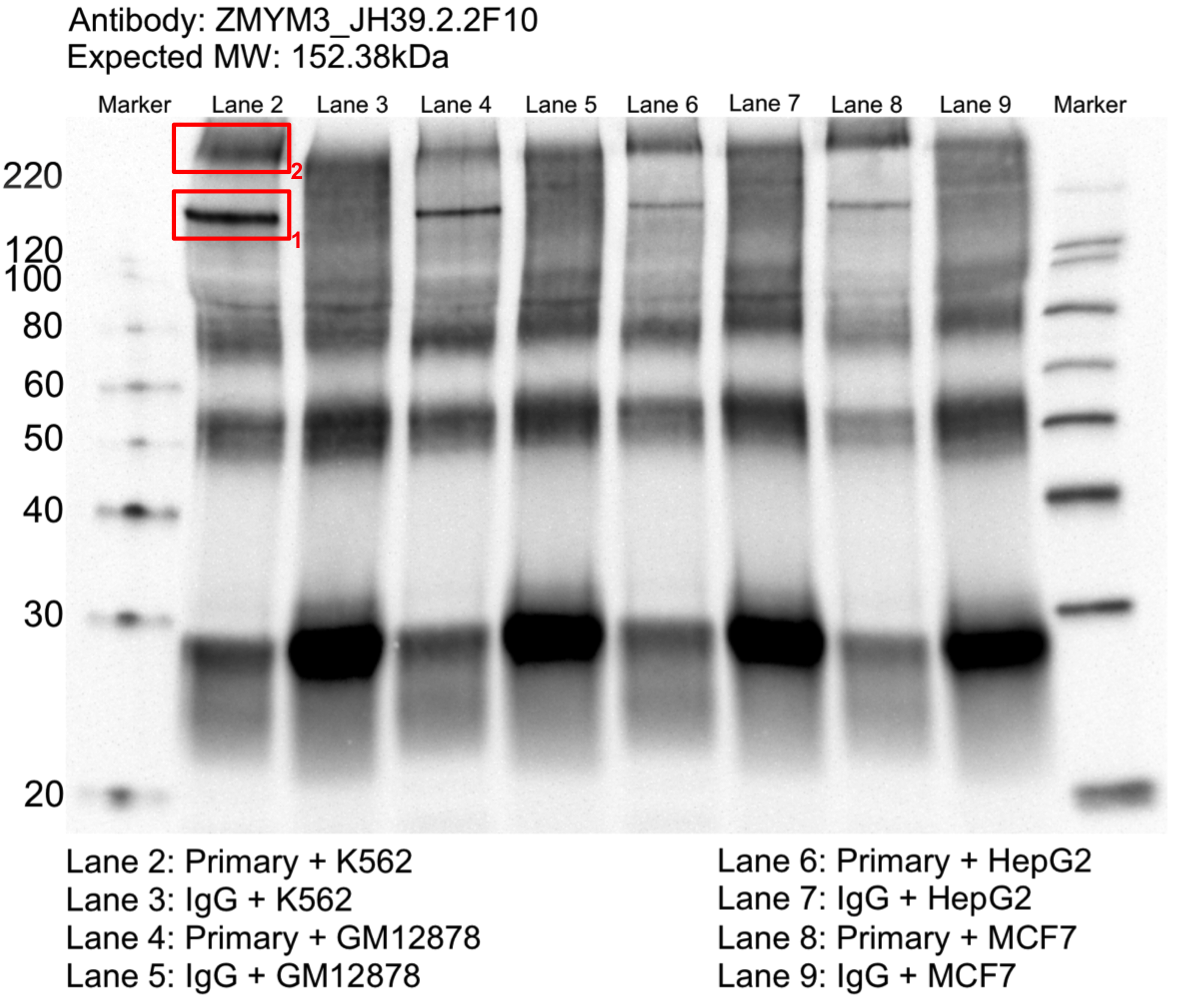
- Whole cell lysates of K562, MCF7, HepG2 and GM12878 were immunoprecipitated using the primary antibody (CDI; JH39.2.2F10). The IP fraction was separated on a 12% acrylamide gel with the Bio-Rad PROTEAN II xi system. After separation, the samples were transferred to a nitrocellulose membrane with an Invitrogen iBlot system. The membrane was probed with the primary antibody (same as that used for IP) and a secondary HRP-conjugated antibody. The resulting bands were visualized with SuperSignal West Femto Solution (Thermo Scientific). Protein Marker (PM) is labeled in kDa. Two bands were detected at ~150 and 220 kDa. Expected size ~152 kDa.
- Submitter comment
- --
- Reviewer comment
- Rescued by mass spectrometry analysis
- Submitted by
- Mark Mackiewicz
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998
- Download
- ZMYM3_IP-WB.png