ENCBS283TEI / cell line
Summary
- Status
- released
- Term name
- K562
- Term ID
- EFO:0002067
- Summary
- Homo sapiens K562 cell line genetically modified (insertion) using CRISPR targeting H. sapiens PBX2
Attribution
ENCODE3 project
- Lab
- Richard Myers, HAIB
- Award PI
- Richard Myers, HAIB
- Submitted by
- Collin White
- Source
- ATCC
- Project
- ENCODE
- External resources
- Aliases
- richard-myers:PBX2-FLAG-k562-1
Genetic modifications
Accession | Category | Purpose | Method | Nucleic acid delivery method | Site |
|---|---|---|---|---|---|
| ENCGM249WDX | insertion | tagging | CRISPR |
|
Donor information
- Status
- released
- Accession
- ENCDO000AAD
- Aliases
- encode:donor of K562, bradley-bernstein:Donor of K562 cells
- Species
- Homo sapiens
- Life stage
- adult
- Age
- 53 years
- Sex
- female
- Health status
- chronic myelogenous leukemia (CML)
- External resources
- References
Documents
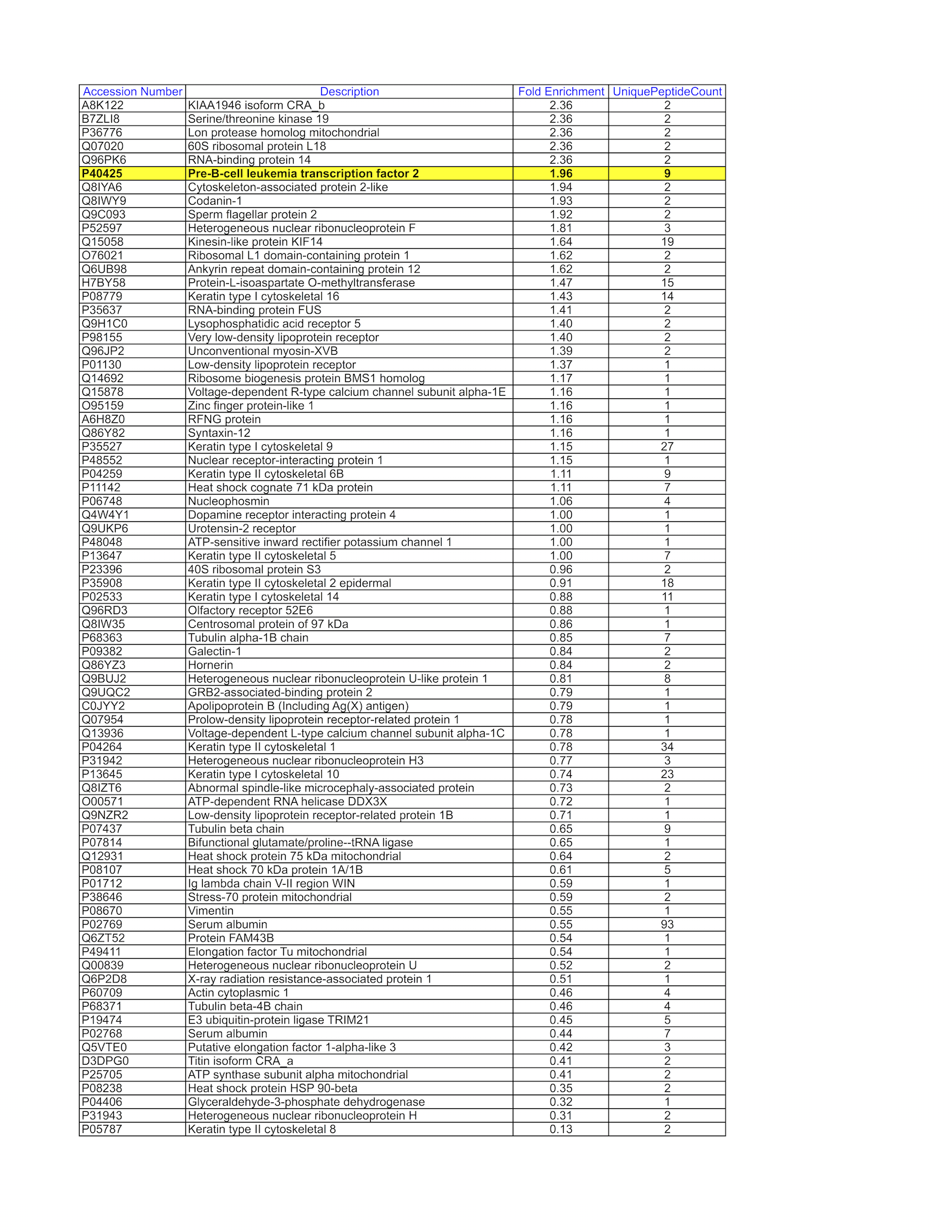
immunoprecipitation followed by mass spectrometry
- Caption
- K562 whole cell lysate was immunoprecipitated using the primary antibody (Sigma; F1804). The IP fraction was loaded on a 12% Bio-Rad TGX gel and separated with the Bio-Rad Tetra Cell system. The whole lane was excised and sent to the University of Alabama at Birmingham Cancer Center Mass Spectrometry/Proteomics Shared Facility. Analysis of whole lane gel from K562: The sample was analyzed on a LTQ XL Linear Ion Trap Mass Spectrometer by LC-ESI-MS/MS. Peptides were identified using SEQUEST tandem mass spectral analysis with probability based matching at p < 0.05. SEQUEST results were reported with ProteinProphet protXML Viewer (TPP v4.4 JETSTREAM) and filtered for a minimum probability of 0.9. All protein hits that met these criteria were reported, including common contaminants. Fold enrichment for each protein reported was determined using a custom script based on the FC-B score calculation from the reference Mellacheruvu et al., 2013. The CRAPome: a contaminant repository for affinity purification mass spectrometry data. Nat. Methods. 10(8):730-736. Doi:10.1038/nmeth.2557. The target protein, ATF1, was identified as the 6th enriched protein and the 1st transcription factor based on IP-Mass Spectrometry.
- Submitted by
- Keenan Graham
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998