ENCAB914RSH
Antibody against Homo sapiens KDM5A
Homo sapiens
K562, HepG2, endothelial cell of umbilical vein, A549
characterized to standards with exemption
Homo sapiens
GM12878
not characterized to standards
- Status
- released
- Source (vendor)
- Abcam
- Product ID
- ab70892
- Lot ID
- GR70517
- Characterized targets
- KDM5A (Homo sapiens)
- Host
- rabbit
- Clonality
- polyclonal
- Antigen description
- lysine (K)-specific demethylase 5A; retinoblastoma-binding protein 2, Jumonji, AT rich interactive domain 1A (RBBP2-like), jumonji AT rich interactive domain 1A
- Aliases
- bradley-bernstein:PchAb 1221
- External resources
Characterizations
KDM5A (Homo sapiens)
Method: ChIP-seq comparison
exempt from standards
- Caption
- This validation relies on the use of a chromatin remodeler and a functionally relevant histone modifier in HepG2 cells, which demonstrate highly similar patterns of enrichment obtained with each antibody. The first track shown used an antibody to H3K4me3 (PchAb 1306, ENCAB848NER), previously characterized to Encode standards, and the second track shown used an antibody to KDM5A (PchAb 1221, ENCAB914RSH).
- Submitter comment
- The Antibody Review Board reviewed this document and the over all experiments to determine the quality of the data. 03-5-20018
- Reviewer comment
- For HepG2, I would support the release of this dataset. The western blot is good, the ChIP-seq correlation between KDM5A and H3K4me3 is good, and the actual data tracks look reasonable. The borderline IDR is due to the fact that this is not a direct DNA binding protein. For GM12878, I would not support the release of the dataset. The ChIP-seq tracks are not good enough (at least to my eye). I note that there was no correlation shown for the data between KDM5A and H3K4me3 in GM12878. I suspect that the correlation would not pass for this dataset.
- Submitted by
- Nina Farrell
- Lab
- Bradley Bernstein, Broad
- Grant
- U54HG006991
- Download
- KDM5A PchAb 1221 SAV.docx.pdf
KDM5A (Homo sapiens)
K562HepG2GM12878endothelial cell of umbilical veinA549
compliant
- Caption
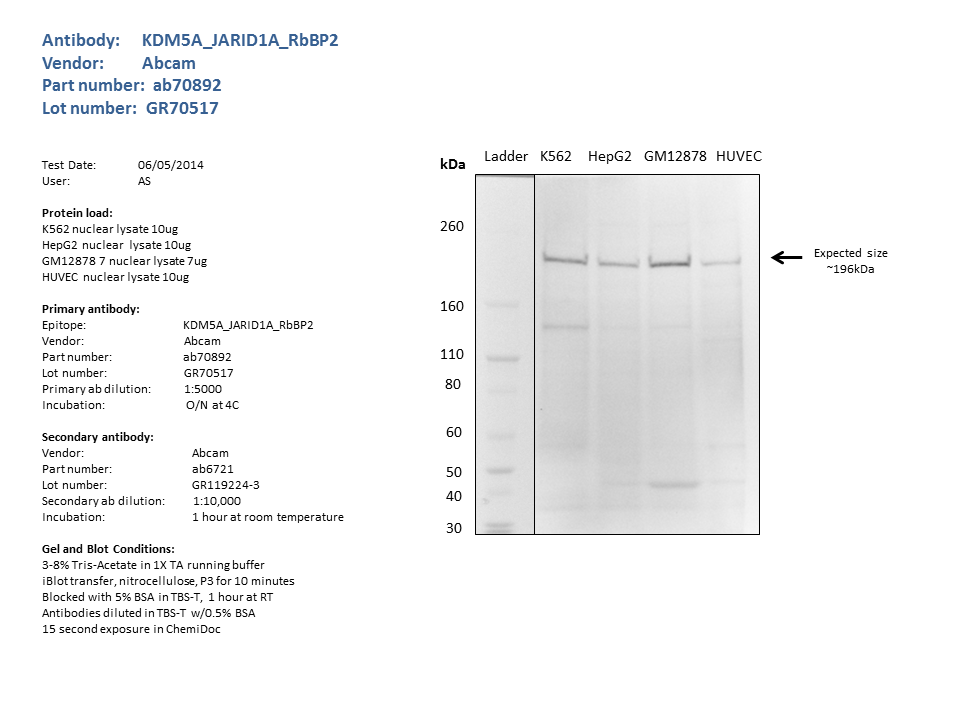
- Nuclear lysates from K562 (10ug), HepG2 (10ug), GM12878 (7ug), HUVEC (10ug), were loaded into a 3-8% Tris Acetate gel in 1X TA running buffer. After separation, the samples were transferred to a nitocellulose membrane using the iblot system. Membrane was blocked for an hour in room temperature, with 5% BSA in TBS-T and blotted with primary antibody in the appropriate concentration over night at 4c. Membrane was washed and blotted with secondary HRP-conjugated antibody. Detection was made with Optiblot ECL Detect Kit (ab133406) for 2 min. a single band of the expected size (~196kDa) was detected.
- Reviewer comment
- Peggy: I would approve exemption of the A549 data . The tracks are convincing and the antibody has been characterized in other cell lines.
- Submitted by
- Noam Shoresh
- Lab
- Bradley Bernstein, Broad
- Grant
- U54HG006991