ENCAB697XQW
Antibody against synthetic tag FLAG, synthetic tag 3xFLAG
Homo sapiens
K562, HepG2, MCF-7, SK-N-SH, WTC11, glutamatergic neuron, HEK293
characterized to standards
- Status
- released
- Source (vendor)
- Sigma
- Product ID
- F1804
- Lot ID
- SLBK1346V
- Characterized targets
- FLAG (synthetic tag), 3xFLAG (synthetic tag)
- Host
- mouse
- Clonality
- monoclonal
- Purification
- affinity
- Isotype
- IgG1
- Antigen description
- Recognizes the FLAG peptide sequence: DYKDDDDK
- External resources
Characterizations
3xFLAG-ZNF124 (Homo sapiens)
compliant
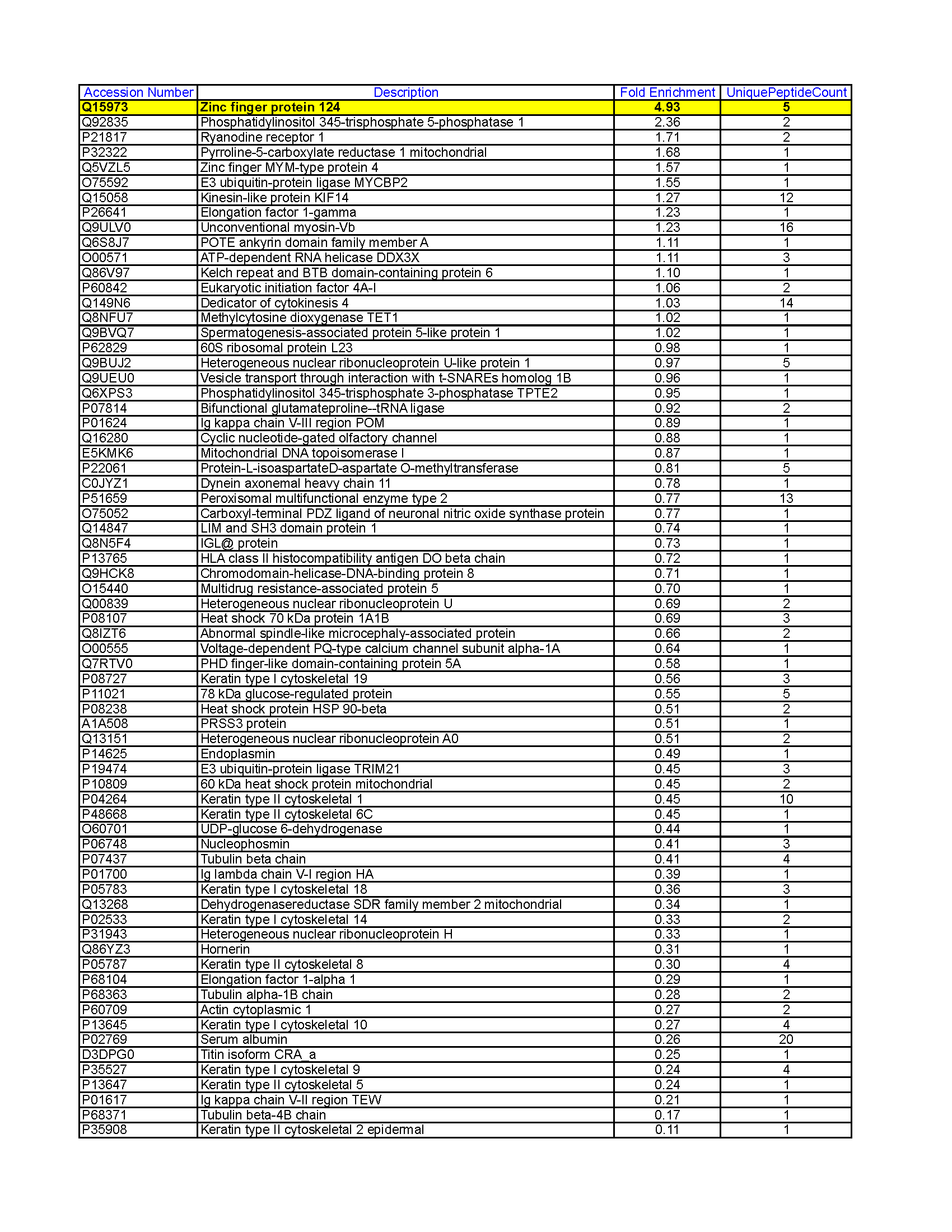
- Caption
- HepG2 whole cell lysate was immunoprecipitated using the primary antibody (Sigma; F1804). The IP fraction was loaded on a 12% Bio-Rad TGX gel and separated with the Bio-Rad Tetra Cell system. The whole lane was excised and sent to the University of Alabama at Birmingham Cancer Center Mass Spectrometry/Proteomics Shared Facility. Analysis of whole lane gel from HepG2: The sample was analyzed on a LTQ XL Linear Ion Trap Mass Spectrometer by LC-ESI-MS/MS. Peptides were identified using SEQUEST tandem mass spectral analysis with probability based matching at p < 0.05. SEQUEST results were reported with ProteinProphet protXML Viewer (TPP v4.4 JETSTREAM) and filtered for a minimum probability of 0.9. All protein hits that met these criteria were reported, including common contaminants. Fold enrichment for each protein reported was determined using a custom script based on the FC-B score calculation from the reference Mellacheruvu et al., 2013. The CRAPome: a contaminant repository for affinity purification mass spectrometry data. Nat. Methods. 10(8):730-736. Doi:10.1038/nmeth.2557. The target protein, ZNF124, was identified as the 1st enriched protein and the 1st transcription factor based on IP-Mass Spectrometry.
- Submitted by
- Mark Mackiewicz
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998
- Download
- FLAG-ZNF124_HepG2_IP-MS.png
FLAG (synthetic tag)
K562HepG2MCF-7
compliant
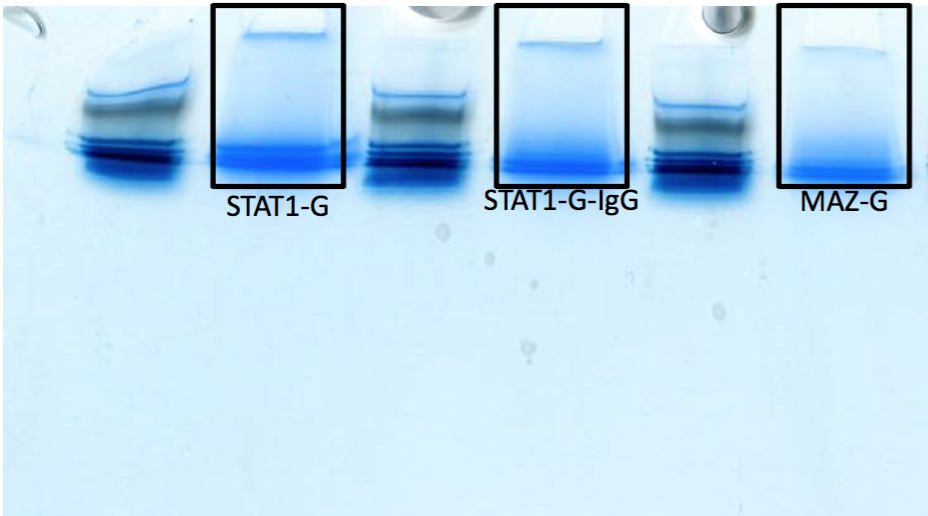
- Caption
- We take the whole lane for IP-MS as opposed to cutting out specific bands. This is an image of what we took as a whole lane for some standard antibody immunoprecipitations. The black boxes represent the fact that we cut the whole lane, we do not run out the gel to get specific bands.
- Submitter comment
- We believe that the mass spec is enough to characterize this antibody and would apply the mass spec rescue scenario described in the standards document.
- Submitted by
- Mark Mackiewicz
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998