ENCAB362XEV
Antibody against Homo sapiens ZNF316
Homo sapiens
K562
characterized to standards
- Status
- released
- Source (vendor)
- Bethyl Labs
- Product ID
- A303-249A
- Lot ID
- 1
- Characterized targets
- ZNF316 (Homo sapiens)
- Host
- rabbit
- Clonality
- polyclonal
- Purification
- affinity
- Aliases
- michael-snyder:AS-1406
- External resources
Characterizations
ZNF316 (Homo sapiens)
K562
not compliant
- Caption
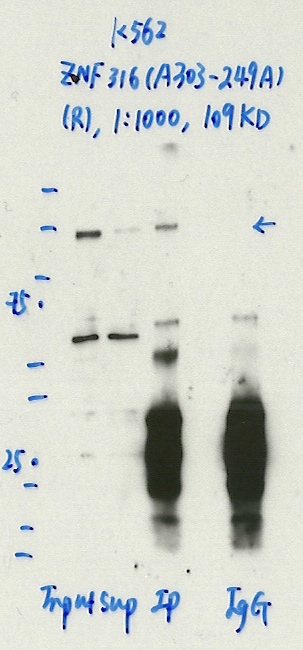
- Immunoprecipitation was performed on nuclear extracts from the cell line: K562, using the antibody A303-249A. The blot shows western blot analysis of input, flowthrough, immunoprecipitate and mock immunoprecipitate using IgG.Molecular Weight: 108.437
- Reviewer comment
- Not within 20% of expected size.
- Submitted by
- Denis Salins
- Lab
- Michael Snyder, Stanford
- Grant
- U54HG006996
- Download
- 1073_09_ZNF316_A303-249A.jpg
ZNF316 (Homo sapiens)
K562
compliant
- Caption
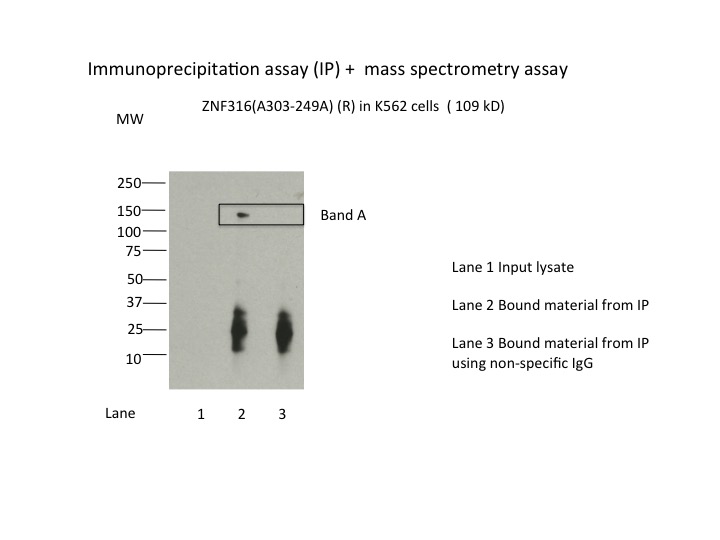
- Immunoprecipitation was performed on nuclear extracts from the cell line K562 using the antibody A303-249A. Lane 1: input nuclear lysate. Lane 2: material immunoprecipitated with antibody. Lane 3: material immunoprecipitated using control IgG. Marked bands were excised from gel and subjected to analysis by mass spectrometry. Target molecular weight: 108.437.
- Submitted by
- Nathaniel Watson
- Lab
- Michael Snyder, Stanford
- Grant
- U54HG006996
- Download
- ZNF316_A303-249A.jpg
ZNF316 (Homo sapiens)
Method: immunoprecipitation followed by mass spectrometry
compliant
- Caption
- IP followed by mass spectrometry. Briefly, protein was immunoprecipitated from K562 nuclear cell lysates using the antibody A303-249A, and the IP fraction was loaded on a 10% polyacrylamide gel (NuPAGEBis-Tris Gel) and separated with an Invitrogen NuPAGE electrophoresis system. The gel was stained by ColloidialCoomassie G-250 stain, gel fragments corresponding to the bands indicated were excised. Then proteins were trypsinized using the in-gel digestion method. Digested proteins were analyzed on an Orbitrap Elite mass spectrometer (Thermo Scientific) by the nanoLC-ESI-MS/MS technique. Peptides were identified by the SEQUEST algorithm and filtered with a high confidence threshold (Peptide false discovery rate < 1%, 2 unique peptides per protein minimum, mass error < 10 ppm).
- Submitter comment
- DDX21, SMARCA5, HNRNPUL1, XRCC5, TOP2A, HNRNPM, AHCTF1, PDCD11, DHX15, BAZ1B, HNRNPR and DNTTIP2 were also detected in MS of ZNF316 (ENCAB363ALA). ACIN1, NOP2, PRPF8, SF3B1, SF3B2 are RNA binding proteins. MKI167 involves poly(A) RNA binding and protein C-terminus binding. EFTUD2 is GTP binding and poly(A) RNA binding protein. BSCL2 is mainly located in endoplasmic reticulum membrane, involves lipid metabolism.
- Reviewer comment
- Peptides ranked higher than ZNF316 were also detected in an independent mass-spec analysis using a different antibody against ZNF316 (ENCAB363ALA).
- Submitted by
- Nathaniel Watson
- Lab
- Michael Snyder, Stanford
- Grant
- U54HG006996
- Download
- ZNF316_A303-249A_final.pdf