ENCAB000AIL
Antibody against Homo sapiens MAX, Mus musculus MAX
Mus musculus
CH12.LX, MEL
characterized to standards
Homo sapiens
liver
characterized to standards with exemption
Homo sapiens
K562
not characterized to standards
- Status
- released
- Source (vendor)
- Santa Cruz Biotech
- Product ID
- sc-197
- Lot ID
- J0809
- Characterized targets
- MAX (Homo sapiens), MAX (Mus musculus)
- Host
- rabbit
- Clonality
- polyclonal
- Purification
- affinity
- Isotype
- IgG
- Antigen description
- Epitope mapping at the C-terminus of Max of human origin
- External resources
Characterizations
MAX (Homo sapiens)
Method: motif enrichment
compliant
- Caption
- The motif for target MAX is represented by the attached position weight matrix (PWM) derived from ENCFF488CNO and ENCFF712NPU. Motif enrichment analysis was done by Dr. Zhizhuo Zhang (Broad Institute, Kellis Lab) using known motifs (http://compbio.mit.edu/encode-motifs/) and previously published ChIP-seq data (http://www.broadinstitute.org/~zzhang/motifpipeline/data/TrainSetInfo.txt). The accept probability score of the given transcription factor was calculated using a Bayesian approach. This analysis also includes three motif enrichment scores, computed by overlapping the motif instances with the given ChIP-seq peak locations. For more information on the underlying statistical methods, please see the attached document. From ENCFF488CNO: Accept probability score: 0.984425787118, Global enrichment Z-score: 1.69262640015, Positional bias Z-score: 4.85974931064, Peak rank bias Z-score: 1.48263455219. From ENCFF712NPU: Accept probability score: 0.980145550336, Global enrichment Z-score: 1.18221297673, Positional bias Z-score: 3.75853018467, Peak rank bias Z-score: 2.07376001231, Enrichment rank: 1.0.
- Submitted by
- Aditi Narayanan
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998
- Download
- ENCAB000AIL.pdf
MAX (Mus musculus)
CH12.LXMEL
compliant
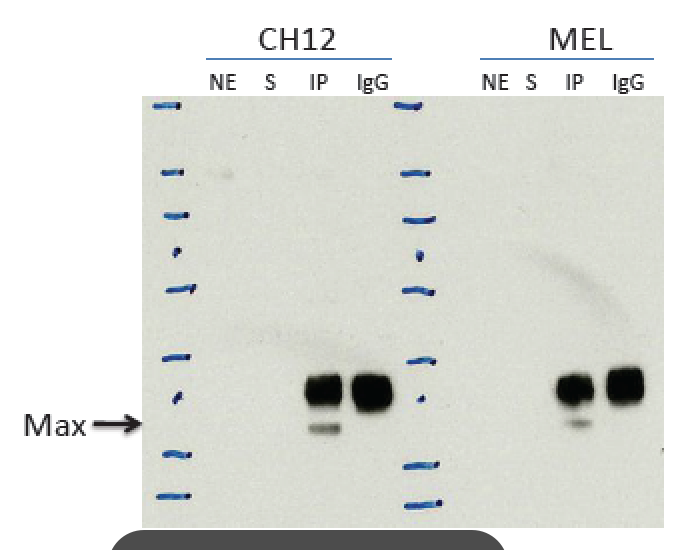
- Caption
- Immunoprecipitation of CH12 and MEL numclear extracts using anti-Max antibody (sc-197) specifically and efficiently enriched a single band of the expected moleculare weight of Max (~21 kD). Arrow indicates immunoprecipitated band of expected size of Max in both CH12 and MEL cell lines (~21 kDa). Second antibody used light chain specific, so only one antibody band. NE: nuclear extract S: supernatant after IP IP: IP with tested antibody IgG: IP with control IgG
- Submitted by
- Michael Snyder
- Lab
- Michael Snyder, Stanford
- Grant
- RC2HG005602
MAX (Homo sapiens)
liver
exempt from standards
- Caption
- The ENCODE Binding Working Group finds for some valuable tissues that recreating a primary on well characterized antibodies is not cost effective. Therefore, they allow exemption from standards for these tissues.
- Submitter comment
- The lab is asking for an exemption for liver cells due to the lack of resource to make a primary characterization for them
- Reviewer comment
- Exempted by the Feb 29, 2016 antibody review panel
- Submitted by
- Richard Myers
- Lab
- Richard Myers, HAIB
- Grant
- U54HG006998
- Download
- No_tissue.png
MAX (Homo sapiens)
K562
not compliant
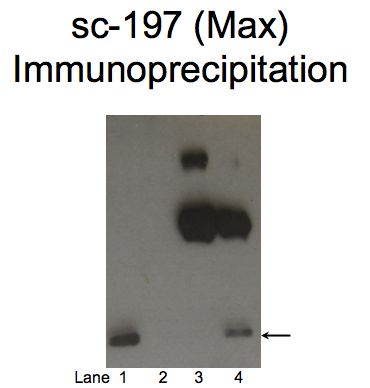
- Caption
- A single band with a mobility consistent with the size expected for Max (19kD; indicated by arrow) is specifically immunoprecipitated from nuclear lysates from K562 cells. Therefore, sc-197 meets this criterion for validation. Immunoprecipitation of Max from K562 cells using sc-197. Lane 1: input nuclear lysate, Lane 2: unbound material from immunoprecipitation assay with sc-197 Lane 3: material immunoprecipitated with sc-197, Lane 4: material immunoprecipitated using control IgG.
- Reviewer comment
- No band in correct lane; band appears in the control lane.
- Submitted by
- Michael Snyder
- Lab
- Michael Snyder, Stanford
- Grant
- U54HG004558
- Download
- sc-197 Immunoprecipitation.png
MAX (Homo sapiens)
not reviewed
- Caption
- Calculations were done by Pouya Kheradpour using a collection of known motifs available at http://www.broadinstitute.org/%7Epouyak/motif-disc/human/. Table 1 shows the fold-enrichments, enrichment p-values and fraction of peaks which contain the motif. The motif which produced the largest value for each criterion is shown in Table 1. Note that while the maximally enriched motifs may differ from the motif with the highest enrichment p-values and the most represented motif, the motifs are highly similar (Figure 1) and thus all values are similar between motifs. Motifs were identified using a matching stringency corresponding to 4-6 (6-mer). Peaks identified by IDR (1% cutoff) were used in the analysis and +/-50bp from peak centers were considered. Enrichments are for a given motif vs. a background consisting of +/- 50bp from the centers of all DnaseI hypersensitive peaks. Repeat mask/simple repeats from UCSC and all gencode v7 exons (including non-protein coding genes) were excluded from the analysis. Comparison to shuffle motifs were used to correct for compositional bias. Enrichment is the corrected # of motifs in ChIP peaks/corrected # of motifs in DNaseI peaks. The current ENCODE standard calls for >4-fold enrichment and >10% motif representation for this criteria to be used for validation. The 3 Max datasets presented here exceed these thresholds and the antibody is considered validated.
- Submitted by
- Michael Snyder
- Lab
- Michael Snyder, Stanford
- Grant
- U54HG004558
- Download
- sc-197 Motif Enrichment .png
MAX (Mus musculus)
not reviewed
- Caption
- Calculations were done by Pouya Kheradpour using a collection of known motifs available at http://www.broadinstitute.org/%7Epouyak/motif-disc/human/. Table 1 shows the fold-enrichments, enrichment p-values and fraction of peaks which contain the motif. The motif which produced the largest value for each criterion is shown in Table 1. Note that while the maximally enriched motifs may differ from the motif with the highest enrichment p-values and the most represented motif, the motifs are highly similar (Figure 1) and thus all values are similar between motifs. Motifs were identified using a matching stringency corresponding to 4-6 (6-mer). Peaks identified by IDR (1% cutoff) were used in the analysis and +/-50bp from peak centers were considered. Enrichments are for a given motif vs. a background consisting of +/- 50bp from the centers of all DnaseI hypersensitive peaks. Repeat mask/simple repeats from UCSC and all gencode v7 exons (including non-protein coding genes) were excluded from the analysis. Comparison to shuffle motifs were used to correct for compositional bias. Enrichment is the corrected # of motifs in ChIP peaks/corrected # of motifs in DNaseI peaks. The current ENCODE standard calls for >4-fold enrichment and >10% motif representation for this criteria to be used for validation. The 3 Max datasets presented here exceed these thresholds and the antibody is considered validated.
- Submitted by
- Michael Snyder
- Lab
- Michael Snyder, Stanford
- Grant
- RC2HG005602
- Download
- sc-197 Motif Enrichment .png